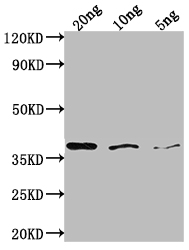
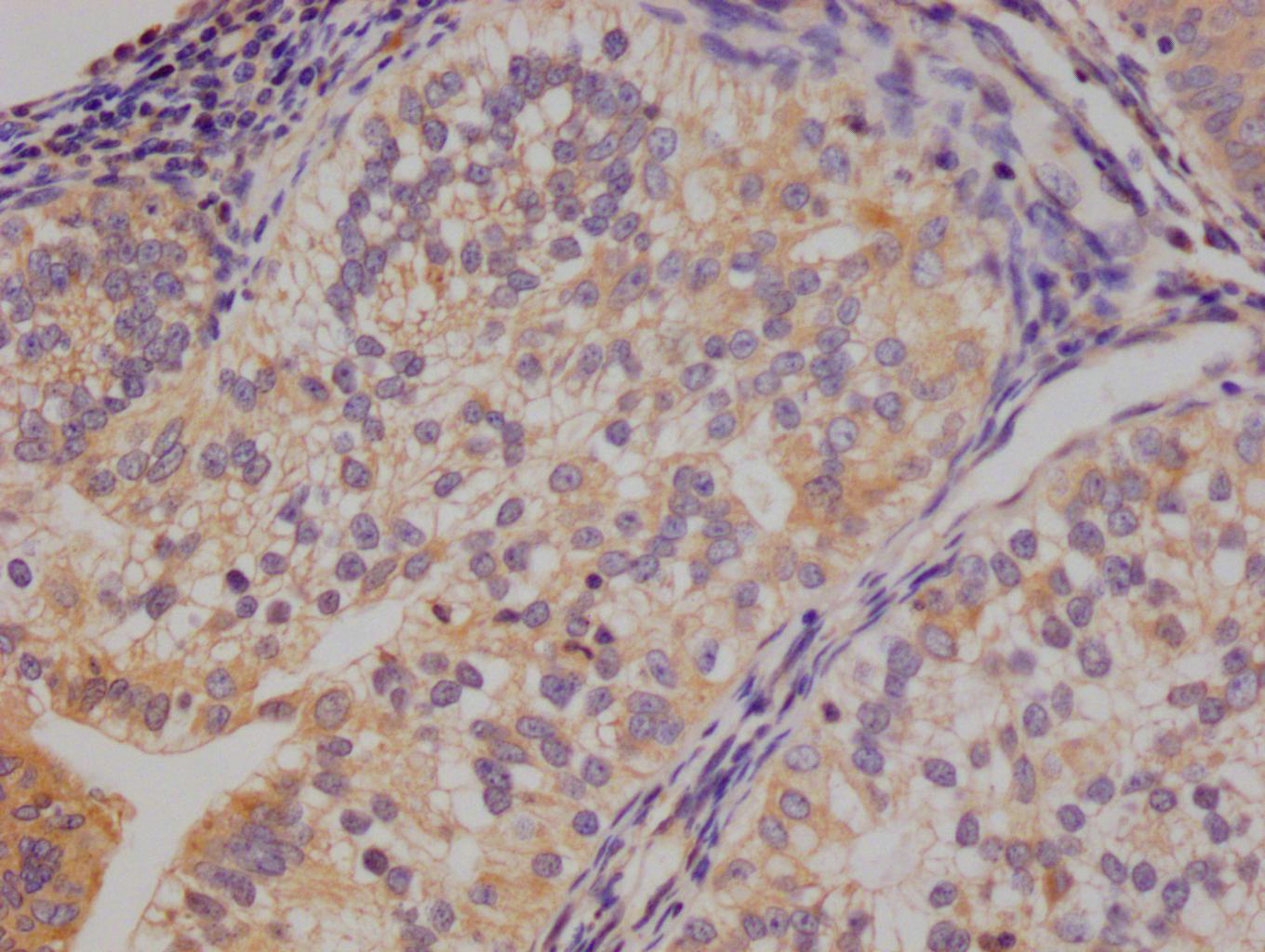
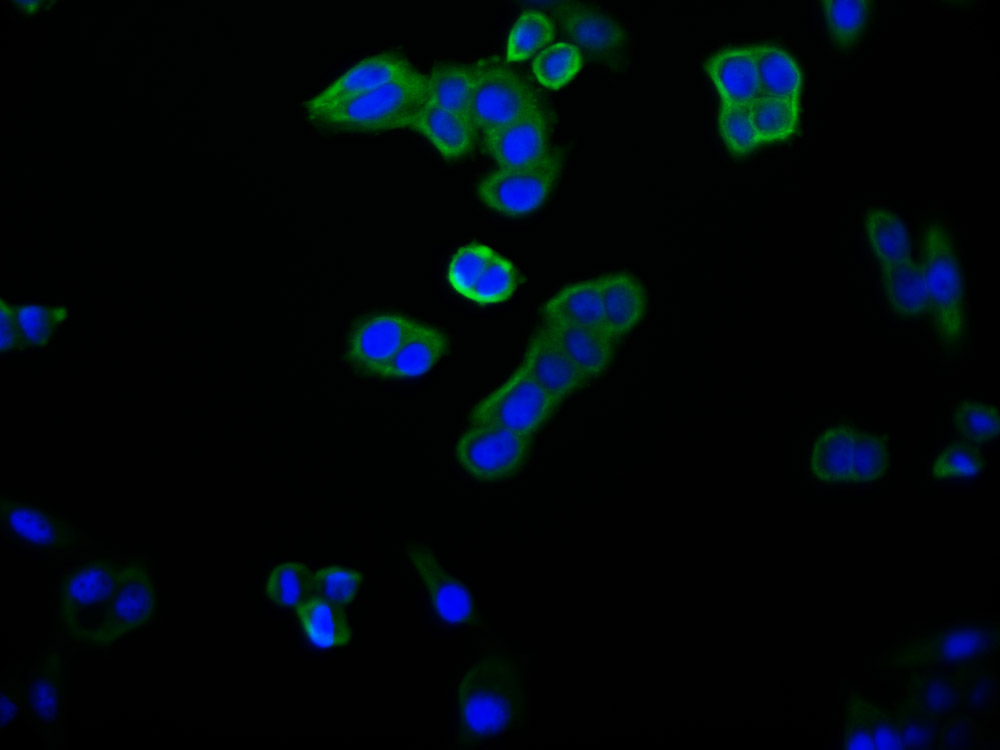
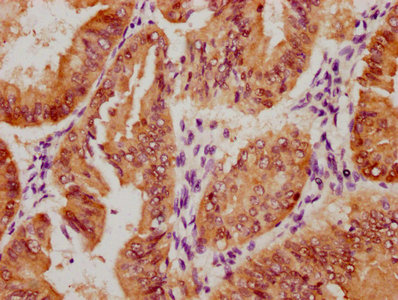
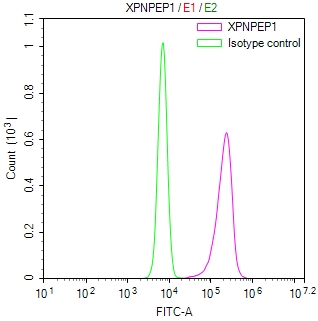
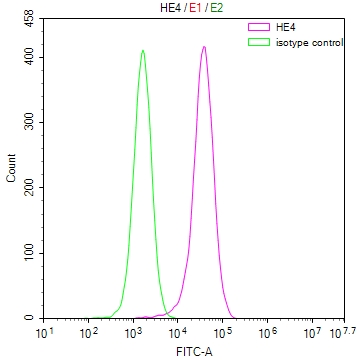
bamA Antibody
-
货号:CSB-PA364271ZA01ENV
-
规格:¥440
-
促销:
-
图片:
-
其他:
产品详情
-
Uniprot No.:P0A940
-
基因名:bamA
-
别名:Outer membrane protein assembly factor BamA (Omp85), bamA, yaeT yzzN yzzY
-
反应种属:Venezuelan equine encephalitis virus (strain Trinidad donkey) (VEEV)
-
免疫原:Recombinant Escherichia coli Outer membrane protein assembly factor BamA (BamA), partial (175-424AA)
-
免疫原种属:Escherichia coli (strain K12)
-
标记方式:Non-conjugated
-
克隆类型:Polyclonal
-
抗体亚型:IgG
-
纯化方式:Protein G
-
浓度:It differs from different batches. Please contact us to confirm it.
-
保存缓冲液:Preservative: 0.03% Proclin 300
Constituents: 50% Glycerol, 0.01M PBS, pH 7.4 -
产品提供形式:Liquid
-
应用范围:ELISA, WB
-
Protocols:
-
储存条件:Upon receipt, store at -20°C or -80°C. Avoid repeated freeze.
-
货期:Basically, we can dispatch the products out in 1-3 working days after receiving your orders. Delivery time maybe differs from different purchasing way or location, please kindly consult your local distributors for specific delivery time.
相关产品
靶点详情
-
功能:Part of the outer membrane protein assembly complex (Bam), which is involved in assembly and insertion of beta-barrel proteins into the outer membrane. Constitutes, with BamD, the core component of the assembly machinery. Efficient substrate folding and insertion into the outer membrane requires all 5 subunits. A lateral gate may open between the first and last strands of the BamA beta-barrel that allows substrate to insert into the outer membrane; comparison of the structures of complete and nearly complete Bam complexes show there is considerable movement of all 5 proteins.; (Microbial infection) Acts as a receptor for CdiA-EC93, the contact-dependent growth inhibition (CDI) effector of E.coli strain EC93; antibodies against extracellular epitopes decrease CDI. Its role in CDI is independent of the other Bam complex components. Is not the receptor for CdiA from E.coli strain 536 / UPEC, which does not have the same mode of toxicity as CdiA from strain EC93; the decreased expression of bamA101 in some experiments decreases the level of outer membrane proteins in general. Susceptibility to CdiA-EC93 is dependent on E.coli BamA; replacing BamA with the gene from S.typhimurium LT2, E.cloacae ATCC 13047 or D.dadantii 3937 renders cells resistant to CdiA-EC93. Cells with BamA from another bacteria no longer form CdiA-EC93-induced aggregates with EC93 cells. A chimera in which E.cloacae extracellular loops 6 and 7 are replaced with loops 6 and 7 from E.coli is susceptible to CdiA-EC93 and to CdiA-CT from strain 536 / UPEC.
-
基因功能参考文献:
- Free-energy calculations of lateral gate opening revealed a significantly lower barrier to opening in the C-terminal kinked conformation; mutagenesis experiments confirm the relevance of C-terminal kinking to BamA structure and function. PMID: 30087180
- Studied how the Bam complex accelerates folding and insertion through the assembly of a slow-folding beta-barrel substrate, LptD. Results suggest a mechanism in which substrate recruitment by periplasmic BamD regulates extracellular loop interactions to activate BamA for folding and insertion. PMID: 29463713
- we studied the assembly of an essential beta-barrel substrate for the Bam complex, BamA. By mutating conserved residues in the beta-barrel domain of this protein, we generated three assembly-defective BamA substrates that stall early in the folding process in the periplasm. Two of the three defective substrates, which harbor mutations within beta-strands, fail to associate productively with the Bam complex PMID: 28223520
- Data show that LptD is folded around LptE, and both components interact with the two essential beta-barrel assembly machine (Bam) components, BamA and BamD. PMID: 27439868
- The s show that electrostatic interactions between the two essential proteins BamA and BamD coordinate conformational changes upon binding of unfolded substrate that allow the assembly reaction to proceed. PMID: 28760846
- Study investigated BamA flexibility using molecular dynamics simulations of BamA embedded in a model Escherichia coli outer membrane, reported that POTRA-membrane interactions influence the set of observed conformations PMID: 27332128
- BamD and BamA mutations cause a marked decrease in the levels of multimeric proteins. PMID: 27161117
- BamA alone can repeatedly facilitate the folding of OMPs PMID: 26394056
- Data show that disulfide crosslinks prevented lateral opening and exit pore formation resulting in a loss of outer membrane proteins BamA function, which can be fully rescued by the reductant tris(2-carboxyethyl)phosphine. PMID: 24980798
- structural analysis of BamB bound to a periplasmic domain fragment of BamA, the central component of the beta-barrel assembly machine PMID: 25468906
- BamA has five polypeptide transport-associated domains in its N-terminus, all of which are essential for BamA protein function. PMID: 24376817
- study determined the crystal structure of the C-terminal transmembrane domain of BamA at 2.6 A resolution; findings suggest the dome over the barrel may play an important role in maintaining the efficiency of OMP biogenesis PMID: 24619089
- These results suggest that the periplasmic region of BamA is firmly attached to the beta-barrel and does not experience fast global motion around the angle between POTRA 2 and 3. PMID: 24530687
- Our direct coupling analysis of BamA implicates residues R661 and D740 in a functional interaction. We find that the substitutions R661G and D740G each confer OM permeability defects and destabilize the BamA b-barrel PMID: 23934888
- Pull-down and Western blotting assays indicate that BamB interacts directly with the POTRA 1-3 domain of BamA PMID: 22948914
- the conserved (641)RGF(643) residues of the BamA L6 loop are important for BamA folding and biogenesis PMID: 22753067
- results imply that BamA and BamD interact directly with outer membrane protein substrates PMID: 22331884
- Taken together, these findings suggest that BamE modulates the conformation of BamA, likely through its interactions with BamD. PMID: 22178970
- Conditions that increase the folding of BamA demonstrate the ability of the reconstituted complex to catalyze more than one round of substrate assembly. PMID: 21823654
- the crystal structure of BamA POTRA4-5 has been determined to 1.50 A resolution with an R factor of 14.7% and an Rfree of 18.9% PMID: 21795783
- The data suggest that SurA and BamA POTRA 1 domain function in concert to assist folding and assembly of most beta-barrel outer membrane proteins except for TolC. PMID: 20598079
- The s demonstrate that (i) BamA is essential for biogenesis of the trimeric auotransporter adhesins YadA, (ii) BamA interacts directly with YadA, (iii) the C-terminal amino acid motif of YadA is important for the BamA-dependent assembly. PMID: 20815824
- the periplasmic chaperone SurA and subunits of the Bam (Omp85) complex catalyse the insertion and assembly of outer-membrane proteins PMID: 19815580
- YaeT may have a role in outer membrane protein assembly in E. coli PMID: 15951436
- YaeT acts as a general outer membrane proteins assembly factor. PMID: 16102012
- YaeT, through its interaction with other outer membrane lipoproteins, forms a functional complex to assemble Escherichia coli membrane protein. PMID: 16824102
- YaeT is composed of two distinct domains, an amino-terminal periplasmic and a carboxy-terminal membrane domain; the periplasmic domain proves to be essential for in vivo function of YaeT PMID: 16829683
- Here we show that an essential and highly conserved gene product, YaeT, is the surface molecule recognized by the majority (ca. 70%) of Stx phages via conserved tail spike proteins PMID: 17693515
- crystal structure of a periplasmic fragment of YaeT reveals the polypeptide transport-associated (POTRA) domain fold and suggests a model for how POTRA domains can bind different peptide sequences PMID: 17702946
- Data show that YaeT-YfgL interaction invloves the region encompassing L173, L175, and R176 of YfgL and it was found that altering residues D227 and D229 in another region of YfgL from E221 to D229 resulted in defective YaeT bindings. PMID: 18165306
- Characterization of mutants resistant to contact-dependent growth inhibition (CDI) allowed the s to identify BamA (YaeT) as the outer membrane receptor for CDI and AcrB as a potential downstream target. PMID: 18761695
- 1H, 13C and 15N chemical shift assignments and secondary structure of the YaeT POTRA domain; a domain found in the Omp85 family of proteins which is critical for insertion and folding of outer membrane proteins in Gram-negative bacteria PMID: 19636842
显示更多
收起更多
-
亚细胞定位:Cell outer membrane.
-
蛋白家族:BamA family
-
数据库链接:
KEGG: ecj:JW0172
STRING: 316385.ECDH10B_0157
Most popular with customers
-
-
YWHAB Recombinant Monoclonal Antibody
Applications: ELISA, WB, IF, FC
Species Reactivity: Human, Mouse, Rat
-
Phospho-YAP1 (S127) Recombinant Monoclonal Antibody
Applications: ELISA, WB, IHC
Species Reactivity: Human
-
-
-
-
-