UNG Antibody
-
货号:CSB-PA025641LA01HU
-
规格:¥440
-
促销:
-
图片:
-
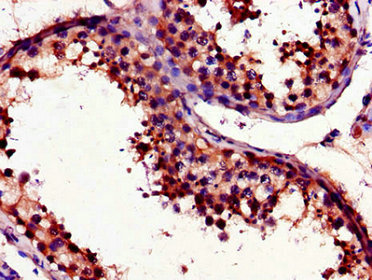
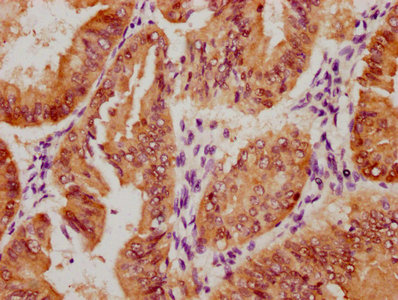
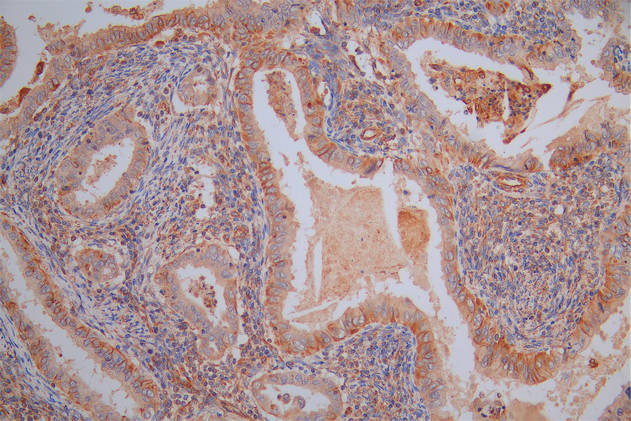
Immunohistochemistry of paraffin-embedded human testis tissue using CSB-PA025641LA01HU at dilution of 1:100
-
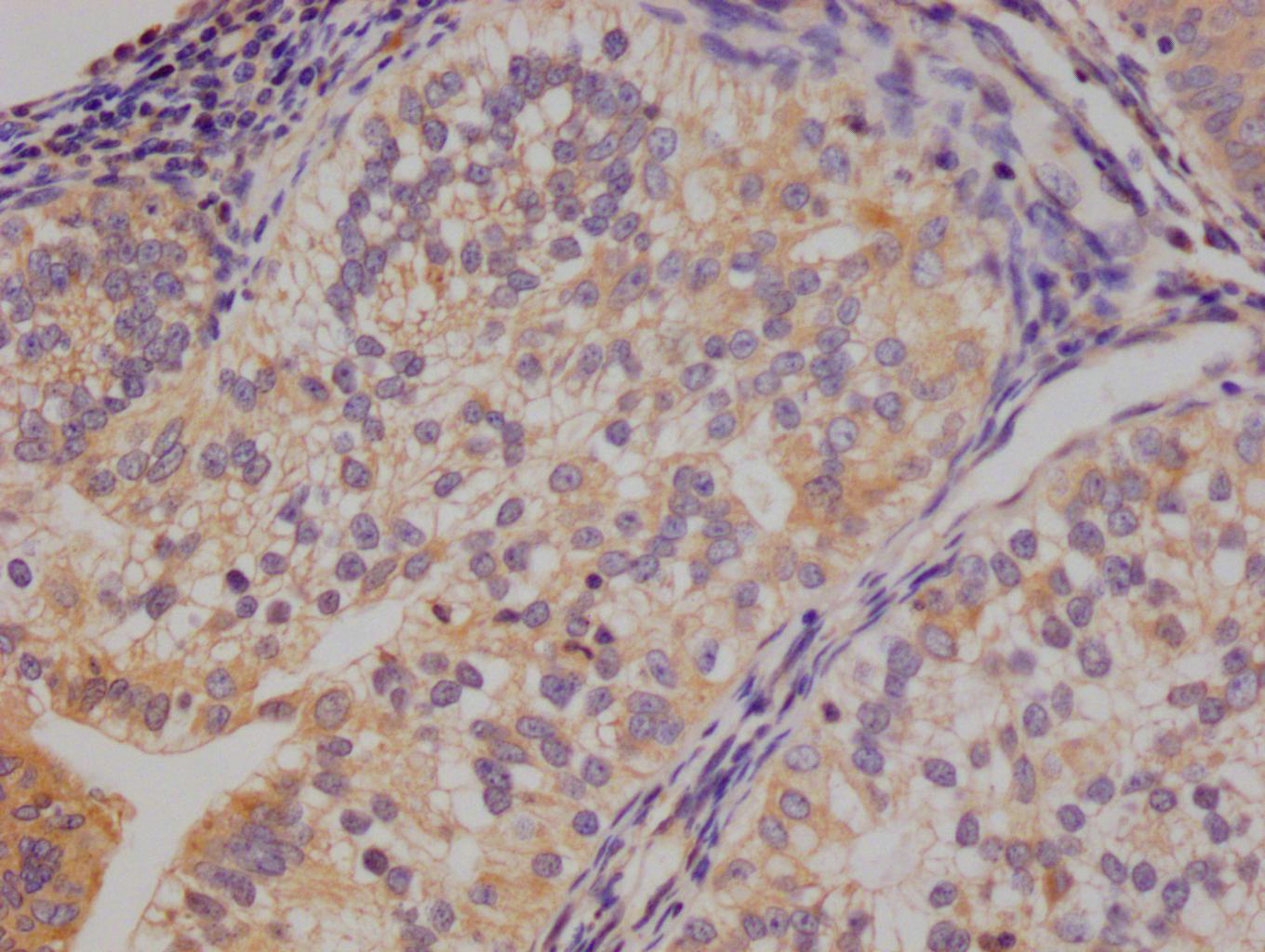
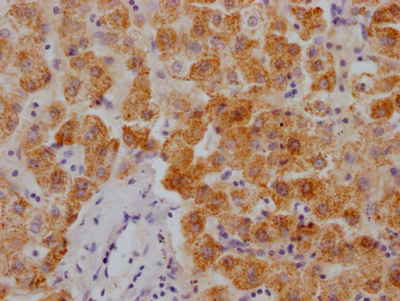
Immunohistochemistry of paraffin-embedded human heart tissue using CSB-PA025641LA01HU at dilution of 1:100
-
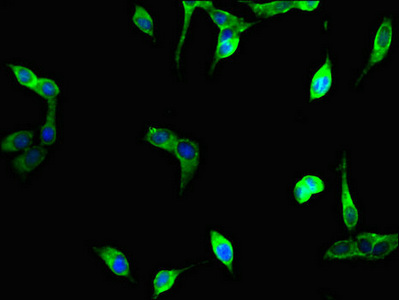
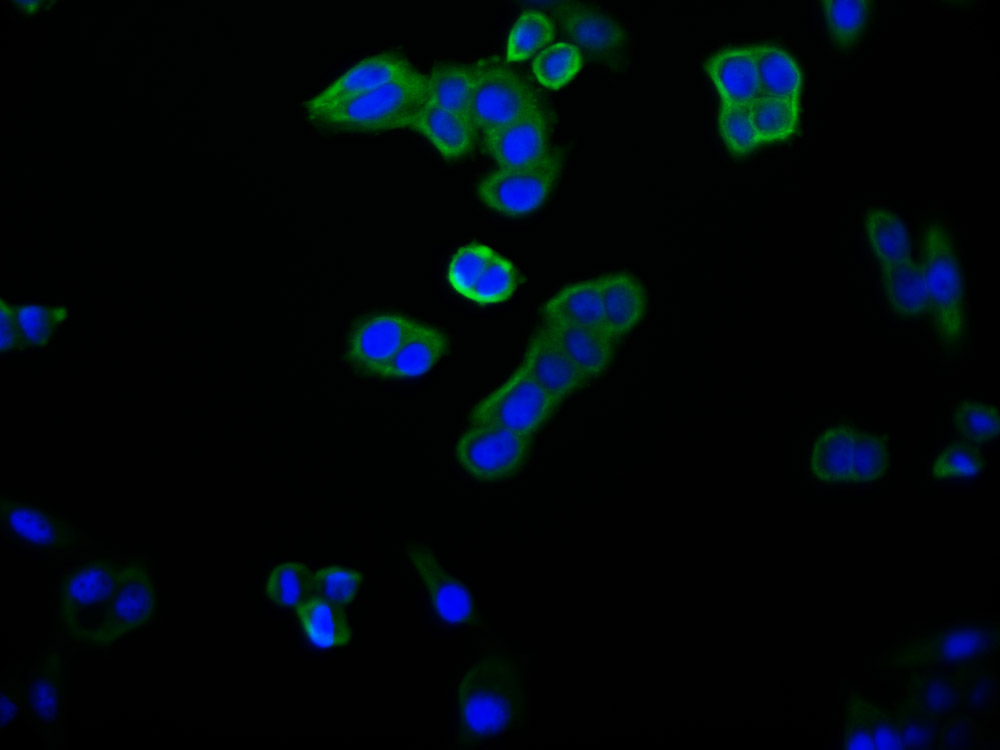
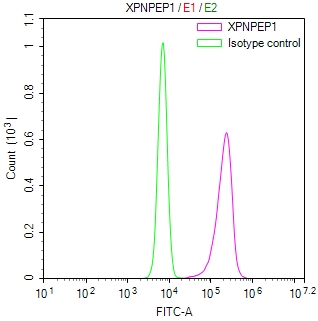
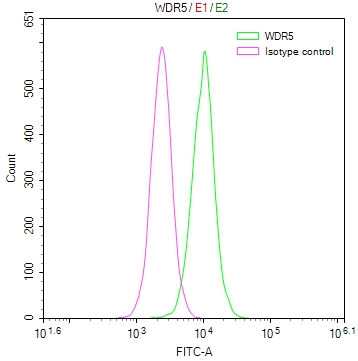
Immunofluorescent analysis of Hela cells using CSB-PA025641LA01HU at dilution of 1:100 and Alexa Fluor 488-congugated AffiniPure Goat Anti-Rabbit IgG(H+L)
-
-
其他:
产品详情
-
产品名称:Rabbit anti-Homo sapiens (Human) UNG Polyclonal antibody
-
Uniprot No.:P13051
-
基因名:
-
别名:DGU antibody; DKFZp781L1143 antibody; HIGM 4 antibody; HIGM4 antibody; OTTHUMP00000240527 antibody; OTTHUMP00000240528 antibody; OTTHUMP00000240529 antibody; UDG antibody; UNG 1 antibody; UNG 15 antibody; ung antibody; UNG_HUMAN antibody; UNG1 antibody; UNG15 antibody; UNG2 antibody; Uracil DNA glycosylase 1 antibody; Uracil DNA glycosylase 2 antibody; Uracil DNA glycosylase antibody; Uracil-DNA glycosylase antibody
-
宿主:Rabbit
-
反应种属:Human
-
免疫原:Recombinant Human Uracil-DNA glycosylase protein (114-224AA)
-
免疫原种属:Homo sapiens (Human)
-
标记方式:Non-conjugated
本页面中的产品,UNG Antibody (CSB-PA025641LA01HU),的标记方式是Non-conjugated。对于UNG Antibody,我们还提供其他标记。见下表:
-
克隆类型:Polyclonal
-
抗体亚型:IgG
-
纯化方式:>95%, Protein G purified
-
浓度:It differs from different batches. Please contact us to confirm it.
-
保存缓冲液:Preservative: 0.03% Proclin 300
Constituents: 50% Glycerol, 0.01M PBS, PH 7.4 -
产品提供形式:Liquid
-
应用范围:ELISA, IHC, IF
-
推荐稀释比:
Application Recommended Dilution IHC 1:20-1:200 IF 1:50-1:200 -
Protocols:
-
储存条件:Upon receipt, store at -20°C or -80°C. Avoid repeated freeze.
-
货期:Basically, we can dispatch the products out in 1-3 working days after receiving your orders. Delivery time maybe differs from different purchasing way or location, please kindly consult your local distributors for specific delivery time.
相关产品
靶点详情
-
功能:Excises uracil residues from the DNA which can arise as a result of misincorporation of dUMP residues by DNA polymerase or due to deamination of cytosine.
-
基因功能参考文献:
- UDG depleted cancer cells displayed sustained DNA damage following 5-FdU treatment PMID: 27517750
- Uracil excision by bacterial UDG and human hUNG2 is reduced at uracils positioned directly 5' or 3' of a G-quadruplex DNA structures. PMID: 26671821
- The UNG1-PRDX3 interaction protected UNG1 from reactive oxygen species-mediated degradation and prevented mitochondrial DNA oxidation. PMID: 27480846
- Characterization of the ability of hUNG2 to translocate on DNA. PMID: 29036472
- Data indicate that DNA glycosylases MYH, UNG2, MPG, NTH1, NEIL1, 2 and 3 on nascent DNA. PMID: 28575236
- Although the N-terminal domain has minimal effects on DNA binding and uracil excision kinetics, this study reports that this domain enhances the ability of human UNG2 to translocate on DNA chains as compared to the catalytic domain alone. PMID: 28787572
- Over-expression of uracil DNA glycosylase 2 in HepG2 protect cells against oxidative stress damage. PMID: 28395718
- Results suggest that the UNG2 N-terminus may serve as a flexible scaffold to tether PCNA and RPA at the replication fork, and that post-translational modifications on the UNG2 N-terminus disrupt formation of the PCNA-UNG2-RPA protein complex. PMID: 28746850
- HIV-1 vpr reprograms CRL4(DCAF1) E3 to direct HLTF for proteasome-dependent degradation independent from previously reported Vpr interactions with base excision repair enzyme uracil DNA glycosylase (UNG2) and crossover junction endonuclease MUS81, which Vpr also directs for degradation via CRL4(DCAF1) E3. PMID: 27335459
- Uracil DNA glycosylase (UNG) is the primary glycosylase responsible for the removal of uracils adjacent to cisplatin ICLs, whereas other uracil glycosylases can process uracils in the context of undamaged DNA. PMID: 28110804
- find that the energetic nature of the observed binding free energies of human 8-oxoguanine DNA glycosylase (hOGG1) and human uracil DNA glycosylase (hUNG) for undamaged DNA are derived from different sources. Although hOGG1 uses primarily nonelectrostatic binding interactions with nonspecific DNA, hUNG uses a salt-dependent electrostatic binding mode PMID: 27571472
- The s show that Vpr can form a trimolecular complex with UNG2 and RPA32 and the positive effect of UNG2 and RPA32 on the reverse transcription process leading to optimal virus replication and dissemination between the primary target cells of HIV-1. PMID: 27068393
- Data suggest novel mechanism in innate immunity allows cytokine TGF-beta to restrict viral circular DNA in hepatocyte nuclei via innate immunity; AID deaminates circular DNA of hepatitis B virus leading to DNA degradation; mechanism depends on UNG. PMID: 26867650
- UNG rs246079 G/A contributes to decreased risk of esophageal cancer in a Chinese population. PMID: 25301111
- base excision by UNG1/2 perturbs transcription of the affected gene PMID: 24951587
- UNG interaction with damaged DNA is dominated by the nonelectrostatic free energy term (DeltaGnon = -7.2 +/- 0.1 kcal mol(-1)), yet retains the nonspecific electrostatic contribution (DeltaGelect = -2.3 +/- 0.2 kcal mol(-1)). PMID: 25408964
- Data shows that cancer cells express UNG2 gene at levels similar to or higher than those seen with peripheral B cells. PMID: 25154417
- UNG specifically interacts with Vpr forming a heterotrimeric complex with DCAF1. PMID: 20870715
- UNG2 colocalizes with CENP-A and H2AX phosphorylation at centromeres in normally cycling cells. PMID: 21399697
- Thus, in experimental models, uracil-DNA glycosylase is a critical mediator of pemetrexed sensitivity that warrants evaluation to determine clinical value PMID: 23873851
- The results obtained suggest the potential role of the g.4235T>C and the c.-31A>G polymorphisms in AMD pathogenesis. PMID: 23714858
- Performed are force-field based molecular dynamics (MD) simulations to explore the structural and dynamical properties of distinct UNG/dsDNA adducts, containing A:U, A:T, G:U, or G:C base pairs, at different stages of the base-flipping pathway. PMID: 23705837
- UNG excises uracil residues from the viral genome during or after cccDNA formation in the nucleus and imply that BER pathway activities decrease the antiviral effect of APOBEC3-mediated hypermutation. PMID: 23696735
- Loss of UNG did not alter cellular sensitivity to 5-FU in two human cell lines. PMID: 23253900
- the search mechanism of uracil DNA glycosylase is not predetermined but, instead, depends on the context in which the uracils are located PMID: 23506270
- the role of the charged DNA phosphate backbone in sliding and hopping by human uracil DNA glycosylase PMID: 23506309
- A comparison of haplotype frequencies between the case and the control revealed that rheumatoid arthritis patients with the Ht2 haplotype are at additional risk for rheumatoid arthritis development PMID: 22138674
- Data show that the R88C variant impairs binding of the R88C variant impairs binding of uracil-DNA glycosylase UNG2 to replication protein A RPA2. PMID: 22521144
- Data show that uracil-DNA glycosylases SMUG1 and UNG2 display widely different sequence preferences. PMID: 22483865
- Excision of AID-induced Uracil by UNG2 occurs predominantly during G1 phase, inducing faithful repair, mutagenic processing, and class switching. PMID: 22529268
- Local enrichment of UNG1 at DNA-bound mtSSB may furthermore facilitate rapid access to- and processing of the damage once the dsDNA conformation is restored. PMID: 22153281
- the first thermal stability analysis of Atlantic cod cUNG and human UNG by differential scanning calorimetry, is reported. PMID: 21959147
- Vpr acts to enhance constitutive DCAF1-dependent UNG2 turnover. PMID: 22292079
- Recruitment of the UNG2 into virus particles increased the infectivity of HIV-1. PMID: 22171270
- measured paramagnetic relaxation enhancements reveal dynamic hot spots in uracil DNA glycosylase that are highly correlated with substrate binding and recognition PMID: 22077282
- Study of the role of histidine-148 provides additional support for inclusion of this amino acid in the list of residues (aspartate-145 and His-268) essential to the chemical step of the UNG2 complete mechanism. PMID: 21473605
- analysis of species specific differences between mouse and humans in regulation of SMUG1 and UNG2 PMID: 21454529
- in the absence of CD40-CD154 interactions, there is a marked reduction in SHM and, specifically, mutations of AICDA-targeted G residues in RGYW motifs along with a decrease in transversions normally related to UNG2 activity. PMID: 19023113
- UNG2 is the major enzyme for removal of deaminated cytosine outside of replication foci in single stranded DNA PMID: 12161446
- More global salt bridges are found in human uracil DNA glycosylase (UDG) and are more stabilizing when compared to cod UDG, possibly playing a role in maintaining overall stability and reducing conformational entropy. PMID: 18004790
- XRCC1 complexes performed complete repair of uracil with higher efficacy than UNG2 complexes. PMID: 20466601
- the Vpr-induced decrease of UNG2 level is mainly related to a transcriptional effect PMID: 19625402
- X4 and R5 HIV-1 have distinct post-entry requirements for uracil DNA glycosylase during infection of primary cells PMID: 20371602
- Data indicate that the activity of UDG likely requires "trapping" transiently exposed states arising from the rotational dynamics of DNA on histones. PMID: 20176960
- The results indicated that there was a correlation between high level of UNG expression and the carcinogenesis of esophageal squamous cell carcinoma. PMID: 15969084
- crystal structure of free and DNA-bound enzyme PMID: 12136137
- Data report the characterization of a functional homologue (ung1) from Schizosaccharomyces pombe of the human uracil-DNA-glycosylase 2 (UNG2) gene. PMID: 12810074
- Nuclear uracil-DNA glycosylase expression was induced in activated B cells. PMID: 12958596
- crystallization reveals example of geometric strain, least atomic motion, and electrophile migration in biological catalysis PMID: 14580190
- Showed that UDG1A protein levels decrease during the S phase of the cell cycle and that it's in the ubiquitin-conjugated form when proteolysis is inhibited by inhibitors of proteosomal dependent protein degradation. PMID: 15084312
显示更多
收起更多
-
相关疾病:Immunodeficiency with hyper-IgM 5 (HIGM5)
-
亚细胞定位:[Isoform 1]: Mitochondrion.; [Isoform 2]: Nucleus.
-
蛋白家族:Uracil-DNA glycosylase (UDG) superfamily, UNG family
-
组织特异性:Isoform 1 is widely expressed with the highest expression in skeletal muscle, heart and testicles. Isoform 2 has the highest expression levels in tissues containing proliferating cells.
-
数据库链接:
HGNC: 12572
OMIM: 191525
KEGG: hsa:7374
STRING: 9606.ENSP00000242576
UniGene: Hs.191334
Most popular with customers
-
-
YWHAB Recombinant Monoclonal Antibody
Applications: ELISA, WB, IF, FC
Species Reactivity: Human, Mouse, Rat
-
Phospho-YAP1 (S127) Recombinant Monoclonal Antibody
Applications: ELISA, WB, IHC
Species Reactivity: Human
-
-
-
-
-