资源|肿瘤数据库汇总(收藏级)
日期:2019-08-22 09:44:20

现如今,随着人们生活方式和环境的改变,恶性肿瘤已经成为疾病死亡病因之一。 传统化疗是对抗癌症的常见方法,但它会攻击全身,造成不必要的副作用,如脱发,恶心和疲劳。
高通量检测技术迅速发展,使得与肿瘤相关的组学数据迅速积累。
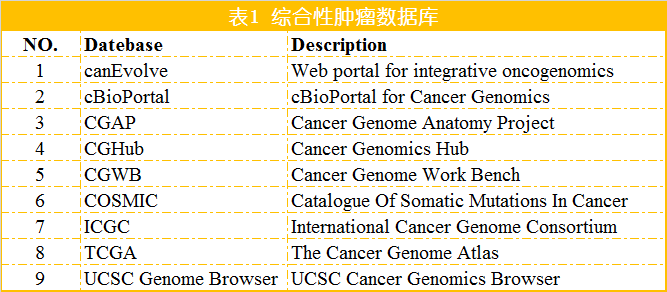
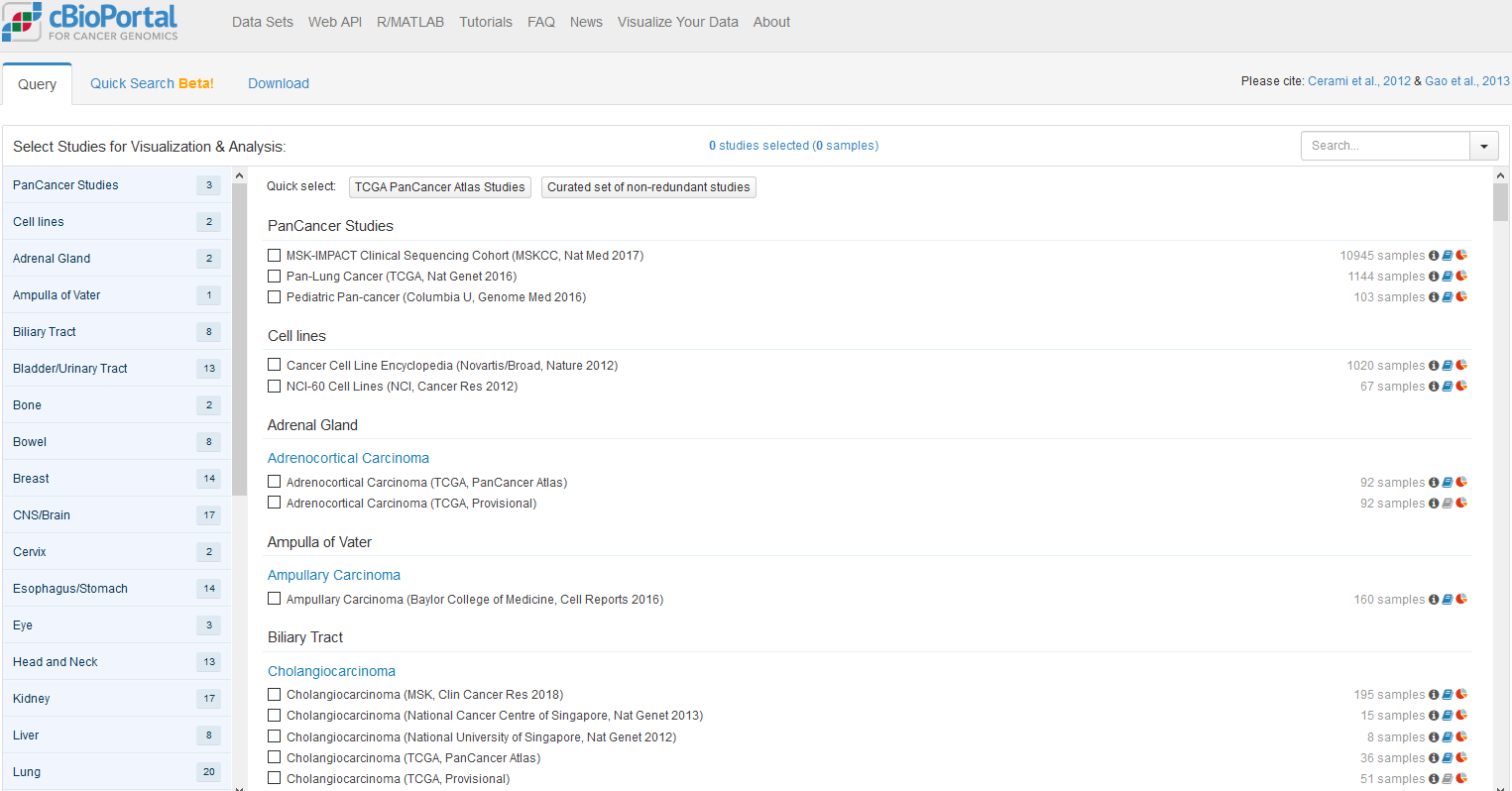
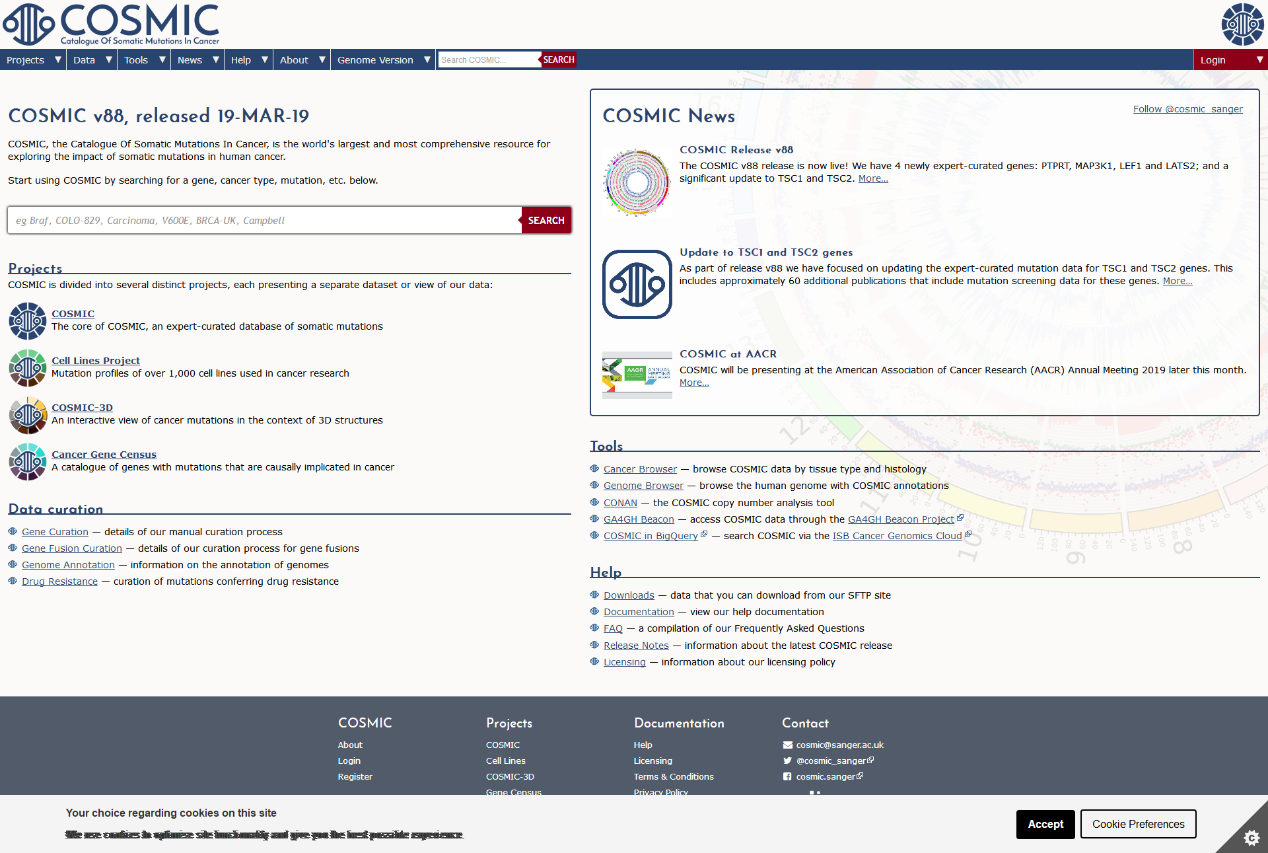

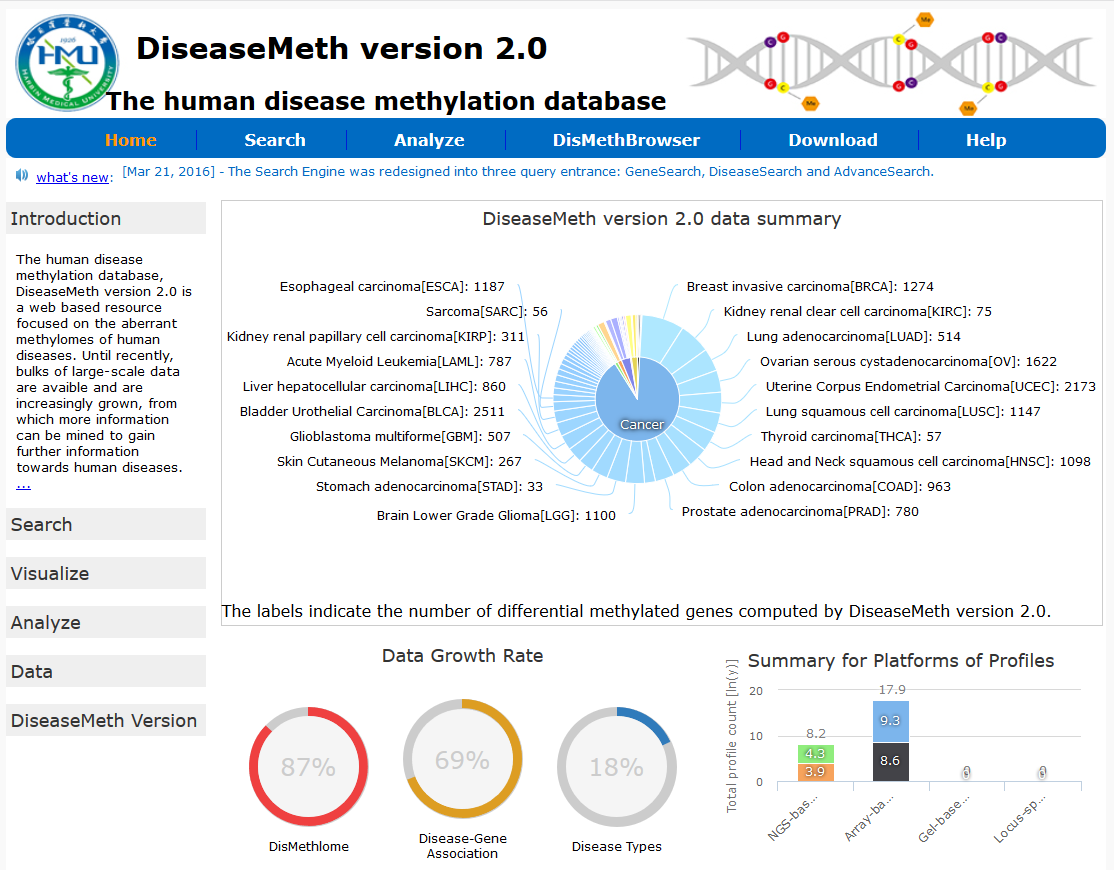
除了上述针对癌症基因组甲基化的数据库外,还有一些数据库搜集和整理更为广泛的甲基化数据,如MethDB和NGSmethDB。 MethDB 是较早的DNA甲基化数据库,主要集中于环境因子对甲基化的影响; NGSmethDB 基于高通量测序数据,最近更新中还包含了SNP信息,以便后续分析。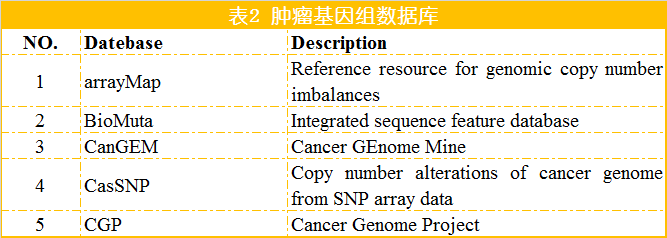 
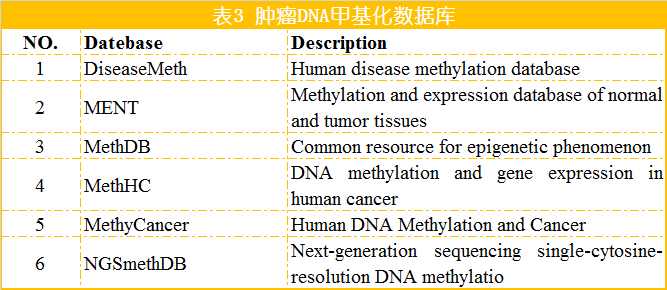

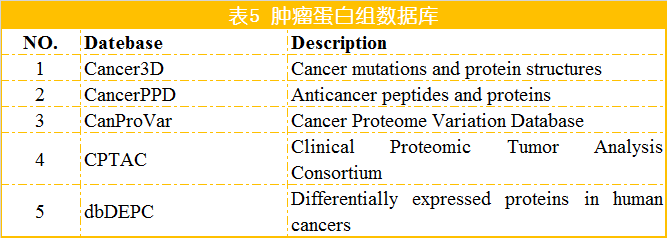
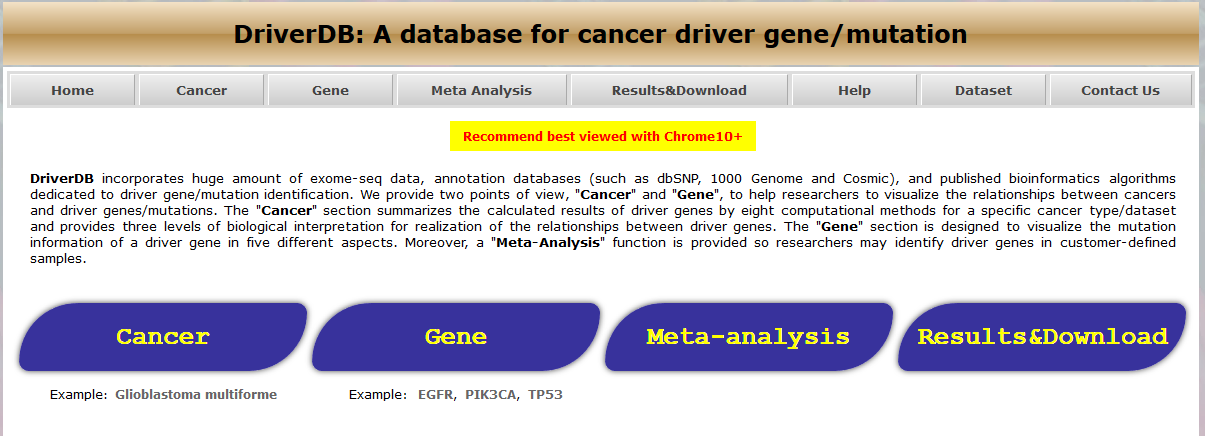
蛋白是生命活动的主要承担者,蛋白结构变异、蛋白修饰的改变以及蛋白含量的变化等导致细胞的生长和代谢变化是肿瘤发生的重要因素。
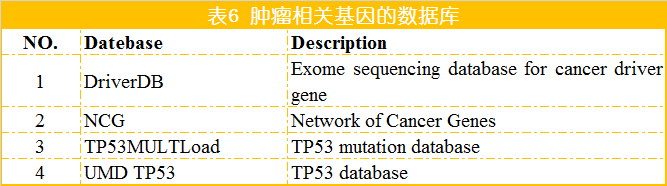
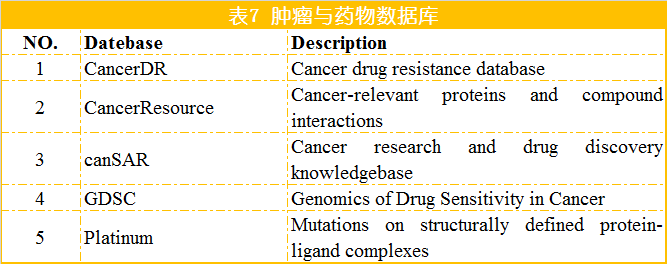


[1] Samur M K, Yan Z, Wang X, et al. canEvolve: A Web Portal for Integrative Oncogenomics [J]. PLOS ONE, 2013, 8. [2] Gao J, Aksoy B A, Dogrusoz U, et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal [J]. Science Signaling, 2013, 6(269): pl1-pl1. [3] Strausberg R L, Buetow K H, Emmert-Buck M R, et al. The Cancer Genome Anatomy Project: building an annotated gene index [J]. Trends in Genetics Tig, 2000, 16(3): 103-106. [4] Wilks C, Cline M S, Weiler E, et al. The Cancer Genomics Hub (CGHub): overcoming cancer through the power of torrential data [J]. Database, 2014. [5] Zhang J, Finney R P, Rowe W, et al. Systematic analysis of genetic alterations in tumors using Cancer Genome WorkBench (CGWB) [J]. Genome Research, 2007, 17(7): 1111-1117. [6] Forbes S A, Beare D, Gunasekaran P, et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer [J]. Nucleic Acids Research, 2015, 43(D1): D805-D811. [7] Banks R, LopezOtín, Carlos. International network of cancer genome projects [J]. Nature, 2010, 464(7291): 993-998. [8] Chang K, Creighton C J, Davis C, et al. The Cancer Genome Atlas Pan-Cancer analysis project [J]. Nature Genetics, 2013, 45(10): 1113-1120. [9] Benz S C, Craft B, Szeto C, et al. The UCSC Cancer Genomics Browser: update 2011 [J]. Nucleic Acids Research, 2013, 43(Database issue): 812-7. [10] Cai H, Gupta S, Rath P, et al. ArrayMap 2014: An updated cancer genome resource [J]. Nucleic Acids Research, 2014, 43(D1). [11] Wu T J, Shamsaddini A, Pan Y, et al. A framework for organizing cancer-related variations from existing databases, publications and NGS data using a High-performance Integrated Virtual Environment (HIVE) [J]. Database, 2014, 2014: bau022-bau022. [12] Scheinin I, Myllykangas S, Borze I, et al. CanGEM: mining gene copy number changes in cancer [J]. Nucleic Acids Research, 2007, 36(Database): D830-D835. [13] Cao Q, Zhou M, Wang X, et al. CaSNP: a database for interrogating copy number alterations of cancer genome from SNP array data [J]. Nucleic Acids Research, 2011, 39(Database issue): D968. [14] Timms B. Cancer genome project to start [J]. European Journal of Cancer, 2000, 36(6): 687. [15] Lv J, Liu H, Su J, et al. DiseaseMeth: a human disease methylation database [J]. Nucleic Acids Research, 2012, 40(Databaseissue): 1030-5. [16] Baek S J, Yang S, Kang T W, et al. MENT: Methylation and expression database of normal and tumor tissues [J]. Gene, 2013, 518(1): 194-200. [17] Huang W Y, Hsu S D, Huang H Y, et al. MethHC: a database of DNA methylation and gene expression in human cancer [J]. Nucleic Acids Research, 2015, 43(D1): D856-D861. [18] He X, Chang S, Zhang J, et al. MethyCancer: the database of human DNA methylation and cancer [J]. Nucleic Acids Research, 2008, 36(Database issue): D836-841. [19] Kolesnikov N, Hastings E, Keays M, et al. ArrayExpress update--simplifying data submissions [J]. Nucleic Acids Research, 2015, 43(D1): D1113-D1116. [20] Frenkel-Morgenstern M, Gorohovski A, Vucenovic D, et al. ChiTaRS 2.1--an improved database of the chimeric transcripts and RNA-seq data with novel sense-antisense chimeric RNA transcripts [J]. Nucleic Acids Research, 2015, 43(D1): D68-D75. [21] Barrett T, Troup D B, Wilhite S E, et al. NCBI GEO: archive for functional genomics data sets - 10years on [J]. Nucleic Acids Research, 2012, 39(D1). [22] Xie B, Ding Q, Han H, et al. miRCancer: a microRNA-cancer association database constructed by text mining on literature [J]. Bioinformatics, 2013, 29(5): 638-644. [23] Rhodes D R, Kalyana-Sundaram S, Mahavisno V, et al. Oncomine 3.0: Genes, Pathways, and Networks in a Collection of 18,000 Cancer Gene Expression Profiles [J]. Neoplasia, 2007, 9(2): 166-180. [24] Wang D, Gu J, Wang T, et al. OncomiRDB: a database for the experimentally verified oncogenic and tumor-suppressive microRNAs [J]. Bioinformatics, 2014, 30(15): 2237-2238. [25] Bhattacharya A, Ziebarth J D, Cui Y. SomamiR: A database for somatic mutations impacting microRNA function in cancer [J]. Nucleic Acids Research, 2012, 41(Database issue). [26] Porta-Pardo E, Hrabe T, Godzik A. Cancer3D: understanding cancer mutations through protein structures [J]. Nucleic Acids Research, 2015, 43(D1): D968-D973. [27] Tyagi A, Tuknait A, Anand P, et al. CancerPPD: a database of anticancer peptides and proteins [J]. Nucleic Acids Research, 2015, 43(D1): D837-D843. [28] Li J, Duncan D T, Zhang B. CanProVar: a human cancer proteome variation database [J]. Human Mutation, 2010, 31(3): 219-228. [29] Ellis M J, Gillette M, Carr S A, et al. Connecting genomic alterations to cancer biology with proteomics: The NCI clinical proteomic tumor analysis consortium [J]. Cancer Discovery, 2013, 3(10): 1108-1112. [30] He Y, Zhang M, Ju Y, et al. dbDEPC 2.0: updated database of differentially expressed proteins in human cancers [J]. Nucleic Acids Research, 2012, 40(D1): D964-D971. [31] An O, Pendino V, D’Antonio M, et al. NCG 4.0: the network of cancer genes in the era of massive mutational screenings of cancer genomes [J]. Database, 2014, 2014: bau015-bau015. [32] Leroy B, Fournier J L, Ishioka C, et al. The TP53 website: an integrative resource centre for the TP53 mutation database and TP53 mutant analysis [J]. Nucleic Acids Research, 2013, 41(Database issue): D962. [33] Kumar R, Chaudhary K, Gupta S, et al. CancerDR: Cancer Drug Resistance Database [J]. Scientific Reports, 2013, 3: 1445. [34] Ahmed J, Meinel T, Dunkel M, et al. CancerResource: a comprehensive database of cancer-relevant proteins and compound interactions supported by experimental knowledge [J]. Nucleic Acids Research, 2011, 39(Database issue): 960-7. [35] Bulusu K C, Tym J E, Coker E A, et al. canSAR: updated cancer research and drug discovery knowledgebase [J]. Nucleic Acids Research, 2014, 42(D1): D1040-D1047. [36] Yang W, Soares J, Greninger P, et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells [J]. Nucleic Acids Research, 2013, 41(Database issue): D955. [37] Pires D E V, Blundell T L, Ascher D B. Platinum: A database of experimentally measured effects of mutations on structurally defined protein-ligand complexes [J]. Nucleic Acids Research, 2014, 43(D1).
肿瘤数据库合辑(附网址)EXCEL免费下载,
关注“武汉j9九游会登录入口首页生物”官方微信,后台回复:
—END—
——j9九游会登录入口首页生物·让科研变得有温度!
必读|神刊CA(IF=244.59)再次发文-2019美国癌症数据速读 必读|影响因子排名第一(IF244.59)神刊CA-2018全球癌症最新统计数据 必读|WHO发布最新预防痴呆症指南-如何早预防? 央视报道|华中地区首个膜蛋白诺贝尔奖工作站落户武汉j9九游会登录入口首页生物 新品推介|高活性EGFR重组蛋白 新品推介|新兴治疗靶点BCMA

上一篇: 哪吒:今天是我生辰宴,都不准哭哦!
下一篇: 资源|七大模式生物数据库











